
Chemoinformatics tools
Institute of Molecular and Translational Medicine, Palacky University
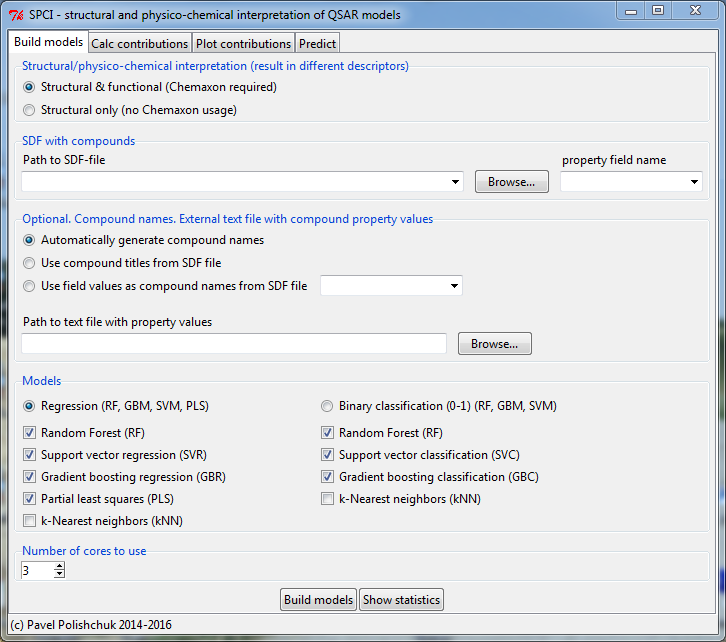
Knowledge-mining software tool for automatic dataset analysis
All you need is a prepared sdf-file with compounds and their property values.
Features:
- Automatic models building, tuning and validation.
- Automatic fragments generation and contribution calculation.
- Plot fragment contributions and save to an external file.
In the framework of development of this project several github repositories were created:
- spci - main module with GUI and all features written in Python
- spci-ext - Python scripts to carry out fragmentation of compounds to further calculate descriptors and make prediction by external software tool and then calculate fragment contributions
- rspci - R package for flexible post-processing and visualization of fragment contributions
Installation and short description of the main software tool is given in the manual
If you found bugs, have questions or propositions please don't hesitate to contact me - pavel_polishchuk@ukr.net
Quick installation notes for Windows users.
To install application just download and unzip into the same folder WinPython and sirms-spci archives.| Installation | OS | Size | Comments |
|---|---|---|---|
| WinPython-32bit-3.5.5.7.zip | Windows | ~360 Mb | WinPython-32bit + all dependencies. Updated 02.05.2015. |
| WinPython-64bit-3.5.5.7.zip | Windows | ~360 Mb | WinPython-64bit + all dependencies. Updated 02.05.2015. |
| sirms-spci.zip | Windows, Linux | ~8 Mb | Main program. Version 0.1.5. Updated 09.05.2016. |
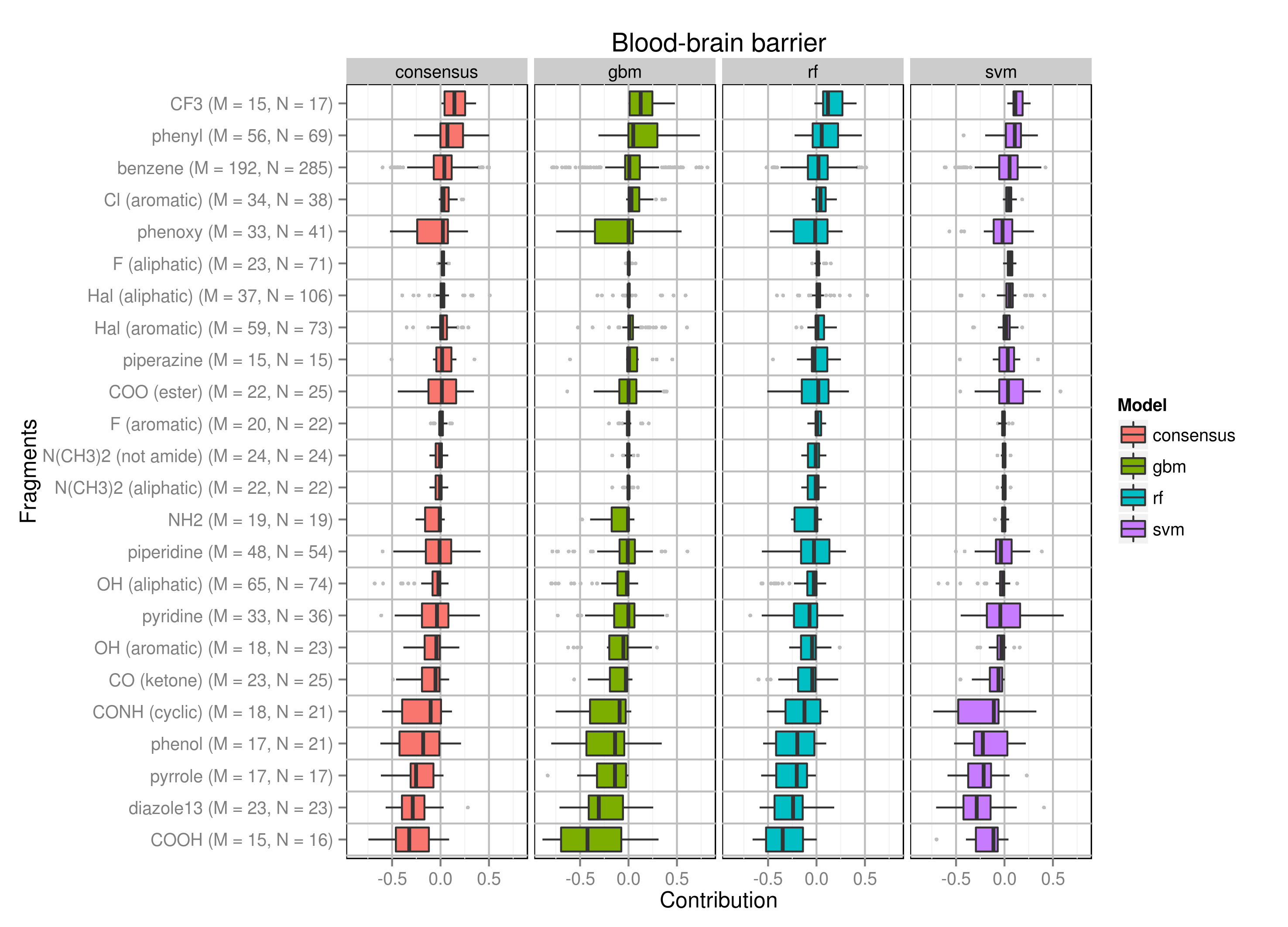
Alternative visualization of output
For those who dislikes Python images and is not familiar with R I recommend to use the developed online tool spci-vis - you just need to upload your text file with calculated fragment contributions and customize plot output. The plot can be saved in different formats.Example output
How to cite
The paper which describes structural interpretation:
P. G. Polishchuk, V. E. Kuz'min, A. G. Artemenko and E. N. Muratov. Universal Approach for Structural Interpretation of QSAR/QSPR Models. Molecular Informatics 2013, 32, 843-853. - http://dx.doi.org/10.1002/minf.201300029
The paper which describes the integrated structural and physicochemical interpretation (SPCI) approach:
Polishchuk P, Tinkov O, Khristova T, Ognichenko L, Kosinskaya A, Varnek A, Kuz’min V. Structural and Physico-Chemical Interpretation (SPCI) of QSAR Models and Its Comparison with Matched Molecular Pair Analysis. Journal of Chemical Information and Modeling 2016, 56, 1455–1469 - http://dx.doi.org/10.1021/acs.jcim.6b00371