
Chemoinformatics tools
Institute of Molecular and Translational Medicine, Palacky University
PMapper - 3D pharmacophore hashes
Encoding of 3D pharmacophores with hashes. This representation is suitable for fast detection of identical pharmacophores without their explicit alignment. It was adapted for 3D ligand-based pharmacophore modeling which requires only 2D strcutures of known active and inactive compounds.
Repositories:
- PMapper - representation of 3D pharmacophores (https://github.com/DrrDom/pmapper)
- PSearch - 3D ligand-based pharmacophore modeling (https://github.com/meddwl/psearch)
- PharMD - MD pharmacophore modeling (https://github.com/ci-lab-cz/pharmd)
The idea is to represent 3D pharmacophore as a complete graph where vertices are pharmacophore features and edges are binned distances between them. All possible quadruplets of features (subgraphs) are considered. For each quadruplet a canonical signature is calculated which encodes topology and stereoconfiguration of a quadruplet. The sorted set of all qudruplets and their counts in a pharmacophore is hashed with MD5 hash to get a 3D pharmacophore hash. Figures below illustrate the procedure, for more details pleases refer our publication.
Generation of 3D pharmacophore hash
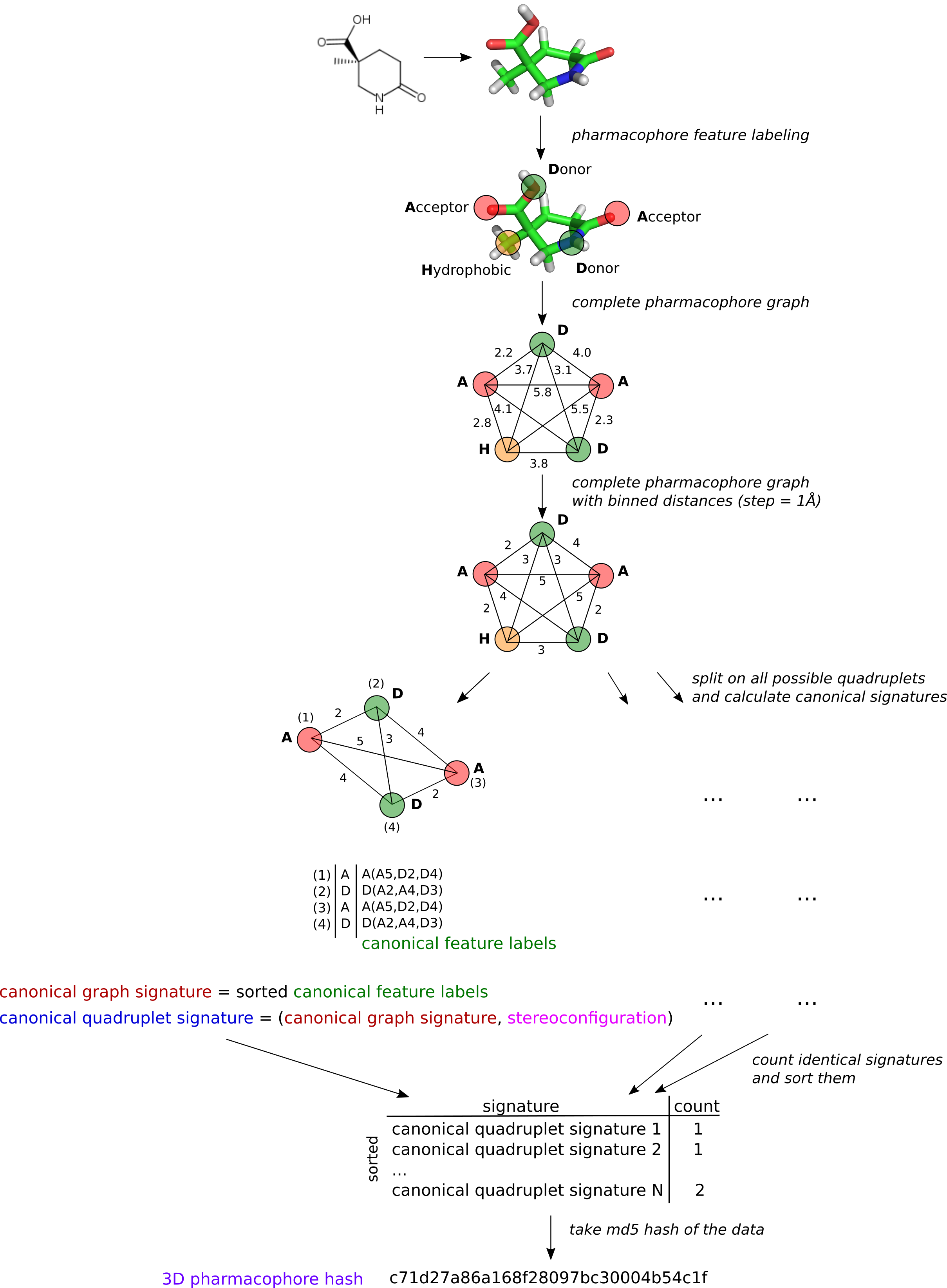
Determination of quadruplet stereoconfiguration

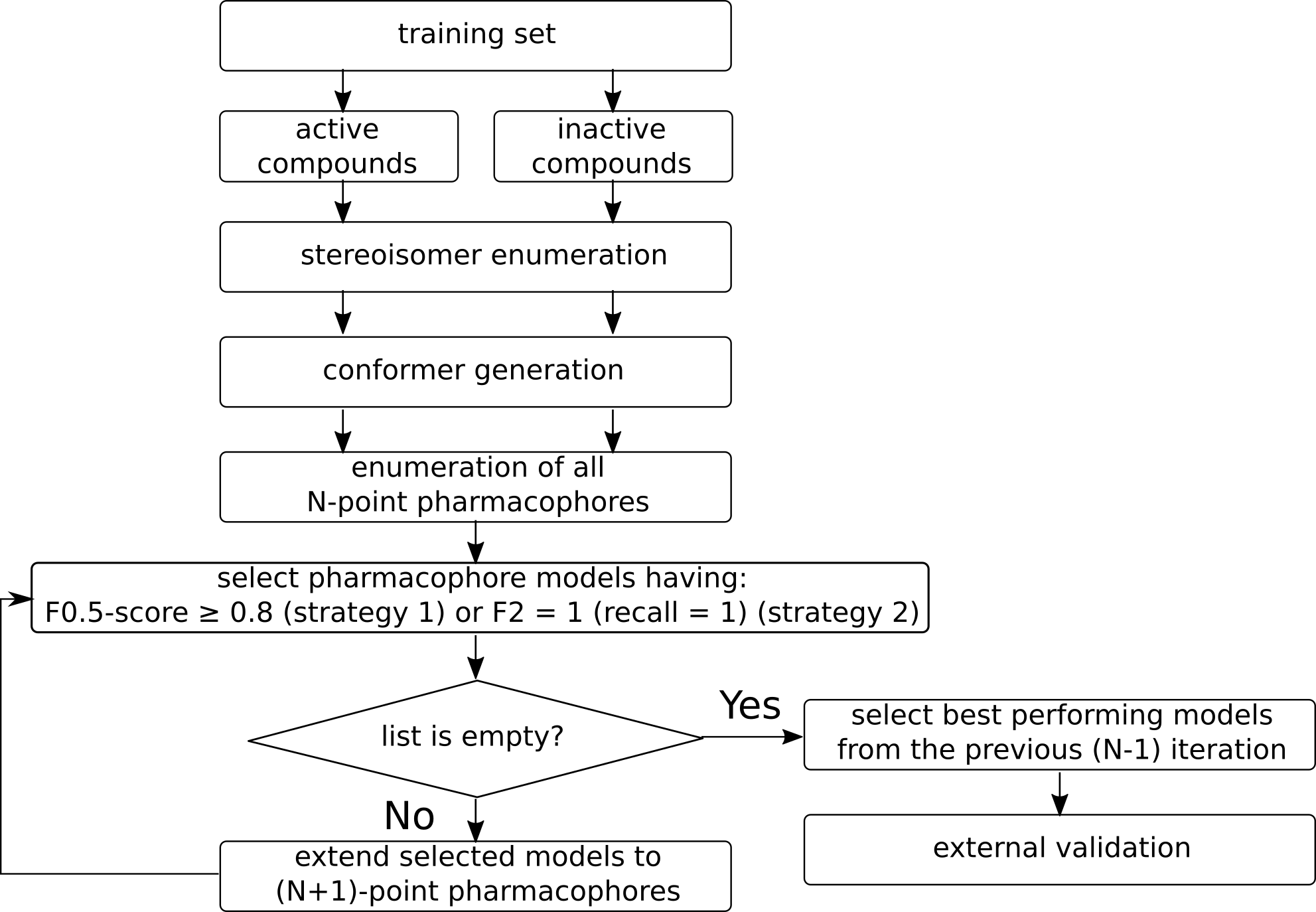
Automated software to generate 3D pharmacophore models based on data sets of known acrive and inactive compounds which does not reqire explicit alignment.
Model generation workflow
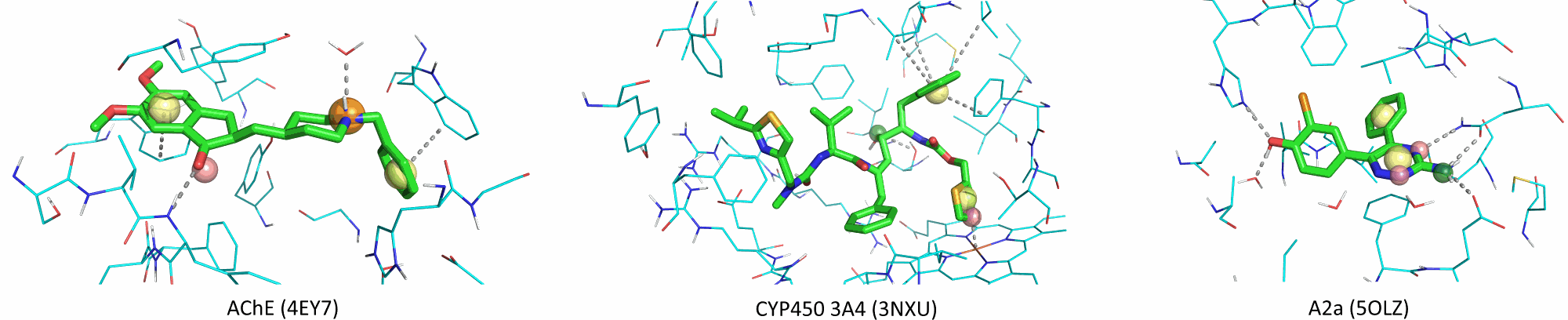
Examples of 3D ligand-based pharmacophores and observed PDB poses
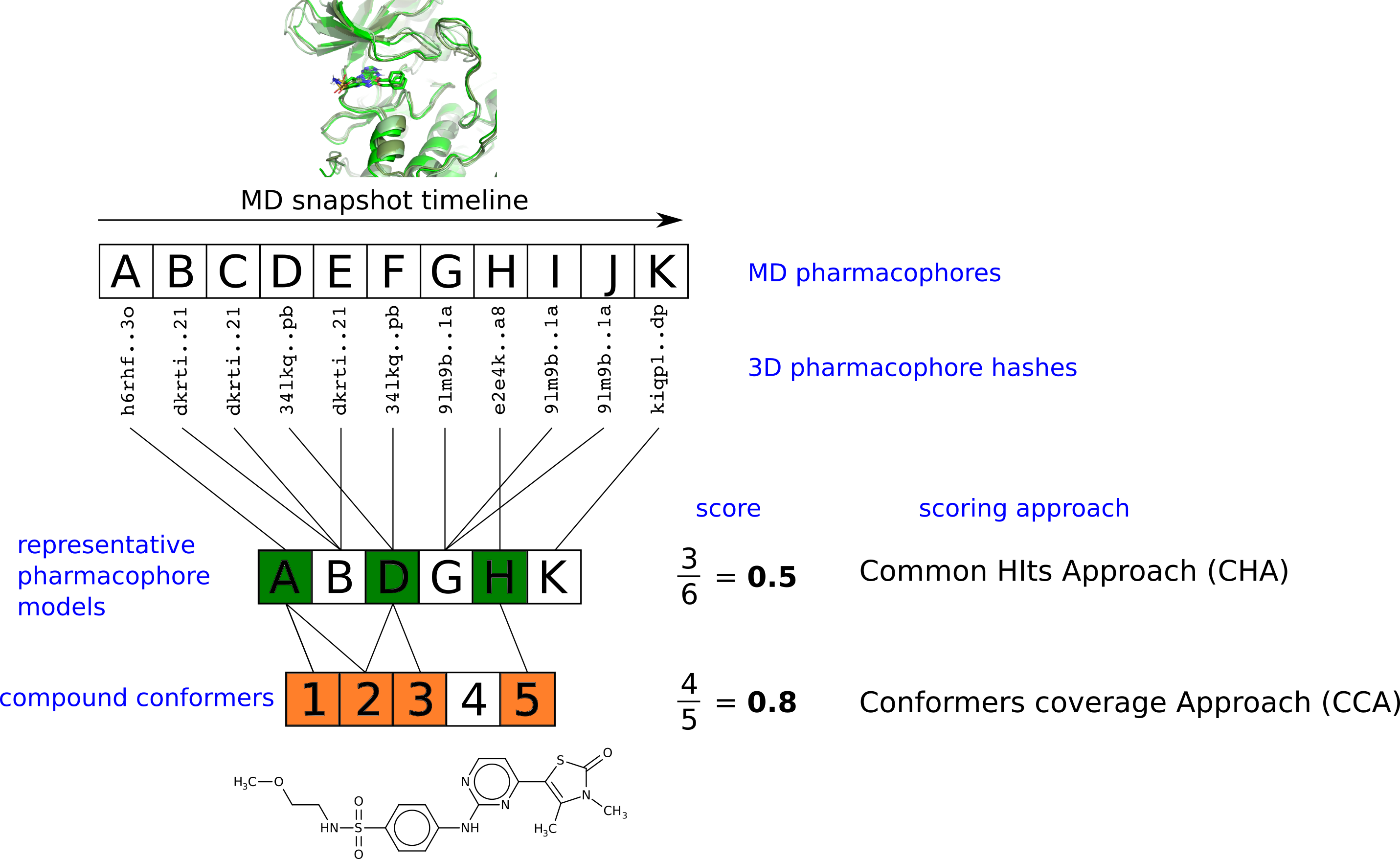
Software to automatically retrieved pharmacophores from MD trajectores of protein-ligand complexes.
Pharmacophore retrieval and ligand scoring workflow
How to cite
The paper which describes 3D pharmacophore representation (PMapper) and 3D ligand-based pharmacohore modeling (PSearch) approaches:
Kutlushina, A.; Khakimova, A.; Madzhidov, T.; Polishchuk, P. Ligand-Based Pharmacophore Modeling Using Novel 3D Pharmacophore Signatures. Molecules 2018, 23, 3094. - https://doi.org/10.3390/molecules23123094
The paper describing MD pharmacophores:
Polishchuk, P.; Kutlushina, A.; Bashirova, D.; Mokshyna, O.; Madzhidov, T., Virtual Screening Using Pharmacophore Models Retrieved from Molecular Dynamic Simulations. International Journal of Molecular Sciences 2019, 20, 5834. - https://doi.org/10.3390/ijms20235834